DryLab®—
Software for Analytical Design Space Modeling
DryLab is the revolutionary HPLC method development and optimization software developed by Lloyd Snyder and his team of fellow researchers. Lloyd's impact on Separation Science is outlined in the following overview.
DryLab predicts chromatograms under a much wider range of experimental conditions than would ever be possible in the laboratory. With DryLab, you can quickly and easily determine exactly how your separation changes as you simultaneously vary multiple method parameters, such as pH, temperature, buffer concentration, and many more. Anybody developing HPLC methods who wishes to optimize complex samples and to economize resources spent developing and running methods will benefit from the many advantages offered by DryLab.
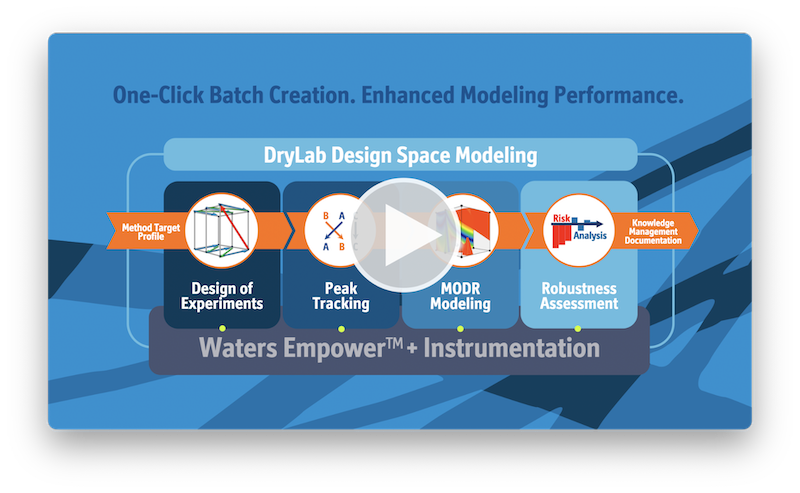
How DryLab works
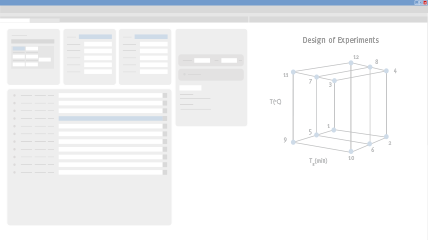
Start by Defining Method Goals, Then Run Input Experiments
The beauty of DryLab is that with data generated from only 2-12 input experiments, DryLab predicts resolution and retention times for millions of unique, virtual chromatograms. The first step to using DryLab is to define your analytical target profile (ATP) and let DryLab help you choose your systematic approach and initial method conditions. Then simply run your input experiments and upload them as *.cdf files, which is the universal format for chromatographic data systems.
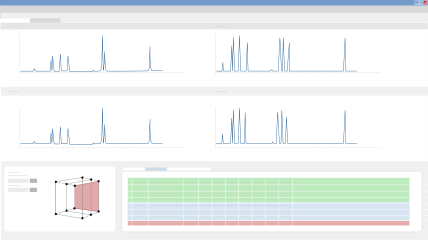
Build Model & Select Working Point
Based on your Design of Experiments (DoE), DryLab creates multi-dimensional resolution models that depict the set of conditions under which you’ll achieve baseline separation or higher. You can view the predicted chromatogram for any point within your model, and with only one mouse click, select the working point with the highest critical resolution. DryLab predicts peak width and retention times with >99% accuracy!
Evaluate Additional Parameters
You can further predict how changes in many additional method parameters affect your separation, such as flow rate, column dimensions, instrument parameters, and eluent in gradient and isocratic conditions. This provides maximum flexibility for method optimization and method transfer.
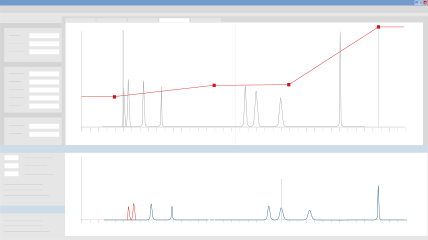
Test Robustness Thresholds
Once you’ve selected your working point, you can test the separation’s tolerance limits by evaluating the impact of small fluctuations on up to six critical separation parameters. You can determine which parameters exert the greatest influence on the method and therefore must be strictly controlled to achieve 100% success in routine applications.
Create GMP Method Documentation
Your last step is to create a comprehensive Knowledge Management report, which pulls all your method data directly from DryLab, and provides you with a platform to justify and comment on your method criteria and choices. This report is GMP compliant and the perfect tool for better knowledge sharing.
Features & Modules
Various features and modules can be added to DryLab Basic:
The identification and assignment of peaks from a set of systematic experiments is an important first step in controlling the HPLC method development process. DryLab’s peak tracking feature includes both peak areas and molecular masses, and offers you an efficient tool for preparing your peak table in an organized and systematic
manner. Within your table you can reorder and turn peak positions, separate double and triple peaks, and reduce complexity. It's color-coded to indicate the likelihood of correct peak identification, and the Comparison Feature checks your original experimental runs against your DryLab model to help further control for errors.
The Gradient Editor is a powerful tool for optimizing your method via gradient elution. While your input experiments must be run with a linear gradient, once you’ve built your DryLab model you can modify the gradient time, change the start and end %B, and add gradient steps. You can
either manually control this process or choose to let DryLab automatically find the best linear or step gradient. This Gradient Editor helps not only to drastically reduce run times, but also to dramatically increase resolution between peak pairs.
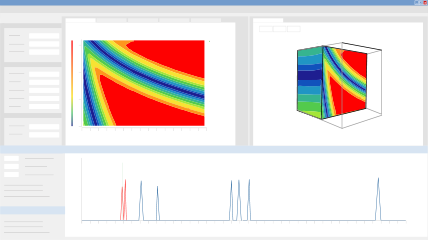
DryLab uses real data to create color-coded maps plotting critical resolution as a function of one or two method parameters. In addition to visualizing the interaction of these parameters, you can also predict chromatograms for changes in other method conditions, such as column dimensions, flow rate, gradient elution, instrument parameters, and many more! Each point within the map corresponds to a unique chromatogram, displayed directly below your resolution map, and you can watch how the resolution changes as you modify the method parameters. Many reversed phase modi, also called Design of Experiments (DoE's) are possible, and you can choose from the following parameters to build your model:
- Gradient Time
- Temperature
- pH
- Ternary Eluent Composition
- Isocratic %B
- Ionic Strength
- Additive Concentration
- Buffer Concentration
By modeling your separation with DryLab, you’ll quickly find optimum separation conditions and gain transparency, understanding, and control of your method.
ColumnMatch lets you compare the selectivity of different columns. Taking into account contributions such as hydrophobicity, steric selectivity, hydrogen-bond-acidity, hydrogen-bond-basicity, and ion-exchange properties of
the silanol groups at different pH-values, ColumnMatch supports you in selecting an equivalent column. In the event that you want to discover hidden peaks, you can also select columns that are very different in their selectivities.
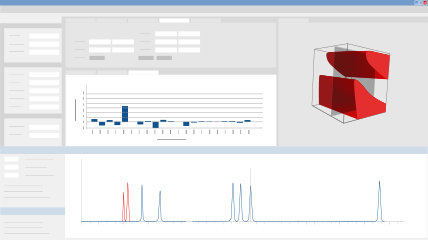
DryLab’s Cube extends the 2D resolution map into the third dimension, providing you with a Method Operable Design Region (MODR) comprised of three parameters in which you can visualize multifactorial variabilities for robust HPLC conditions. In addition, you can model up to 8 other parameters, including column dimensions, flow rate, gradient points, and instrument parameters. The Cube gives you exponentially more conditions in which to model your method, and offers an intuitive display of how simultaneous changes to multiple method parameters affect the critical resolution and selectivity of your separation. A special view shows the 3D regions that fulfill your given resolution requirement (for example, baseline separation of all peaks). You can scroll through the cube to see what the chromatogram looks like for any set of conditions within the 3D region.
Watch the video below to see how resolution and selectivity change as you modify the method conditions: gradient time, temperature, and ternary eluent concentration (%MeOH in %ACN).
The Robustness Module tests the tolerance limits of your selected working point by computing the number of Out-of-Specification (OoS) results that occur given small fluctuations in your method parameters. You can evaluate the following conditions:
- Gradient Time
- Temperature
- pH or Ternary Composition
- Flow Rate
- Start %B
- End %B
A chromatogram is generated for every possible combination of errors and shows you the range of resolution values you can expect during routine application. Based on the number of successful experiments, you may wish to choose a new working point to ensure safer results during routine application (this does not require a new validation). Moreover, you can evaluate which method parameters exert the highest influence on your separation, which is highly useful for setting up an efficient control strategy.
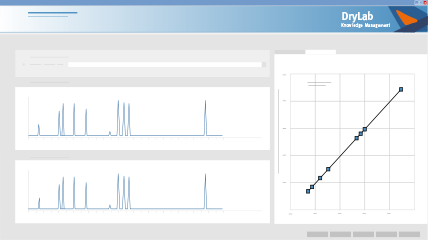
DryLab Knowledge Management is your reporting tool for documenting and archiving your method. It encourages a Quality by Design (QbD) approach to method development and ensures that your method conforms to these standards by providing a comprehensive method report, including a platform for the step-by-step justification of your method choices. By implementing DryLab Knowledge Management, you’ll not only achieve excellent GMP documentation of your method, but also more easily and effectively collaborate between departments, and support
analytical method transfer during development and manufacturing.
DryLab Knowledge Management provides an Analytical Method Summary to be signed and dated by the author and supervisor, making it GMP compliant and the perfect documentation for inspections.
Take a look at these recent Knowledge Management Reports for two UPLC methods: Ebastine and Amlodipine
Creating sample sets needs plenty of experience, time, and scrutiny. Launching the Automation Module, DryLab allows for cost-effective compliance with principles of Analytical Method Lifecycle Management.
DryLab's Automation Module creates a complete SampleSet with one-click based on the input conditions and preferences and takes into consideration equilibration times, blanks, and/or run repetitions.
The DryLab-Empower (OpenLab) connection simplifies complex analytical procedures, improving the reproducibility and accuracy of the results across the LC techniques. It eliminates human error by excluding manual interaction while transferring data between the modeling software and CDS, bringing it onto a daily routine level, hence, yielding a reasonable profit for the companies.
Additionally, DryLab's Automation Module can perform method modeling across multiple columns of different chemistries or several samples at-a-time. A drug manufacturer can reduce the analysis time by switching to a new column.
Please listen to the LCGC podcast linked below to get an overview of workflow advantages and future developments of DryLab's Automation Module.
The Design Space Comparison Module enables users to compare multiple DryLab models (Design Spaces) generated under different conditions, such as varying stationary phases. This powerful feature provides a quick overview of how different columns perform for a specific sample and helps identify common method conditions that are effective across all columns, establishing Column Equivalency.
Aligned with the principles of ICH’s Analytical Quality by Design guideline draft (Q14), this module supports a knowledge-driven approach to analytical procedure development. It emphasizes risk assessment and management to ensure robust and
scientifically sound analytical outcomes. DryLab’s Design Space Comparison and Automation Modules are designed to put these quality-by-design principles into practical use. Recent studies highlight their effectiveness in identifying column interchangeability and in advanced applications such as the analysis of SARS-CoV-2 antibodies using automation workflows.
As Design Space Modeling becomes a cornerstone of modern analytical procedure development, its role is growing, especially for enabling real-time decision-making in the data-driven pharmaceutical industry.